Frontiers | Extensive Variation in Gene Expression is Revealed in 13 Fertility-Related Genes Using RNA-Seq, ISO-Seq, and CAGE-Seq From Brahman Cattle

CAGE-seq characterization of alternative promoter usage and full-length... | Download Scientific Diagram
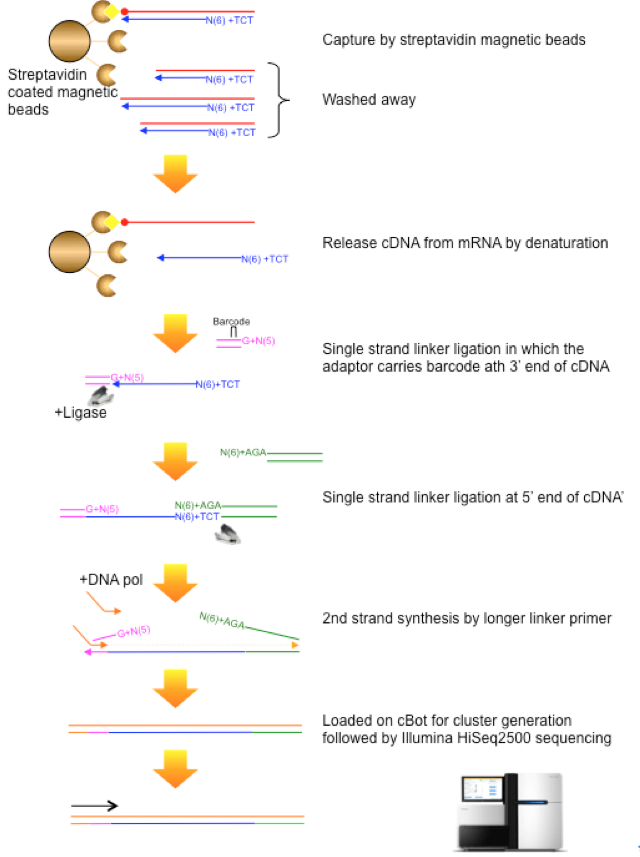
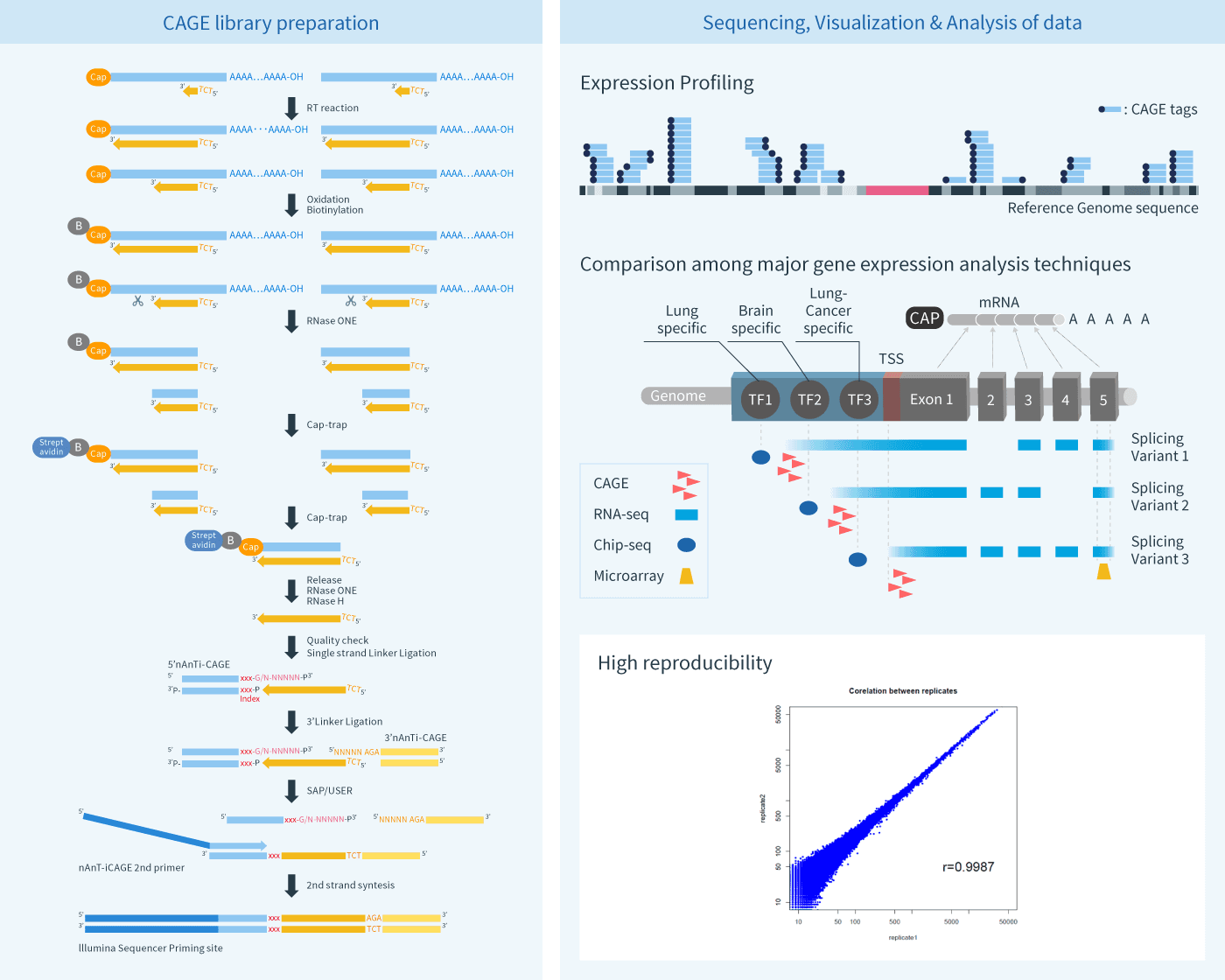
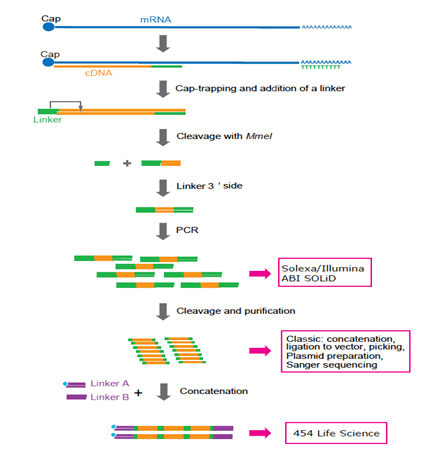
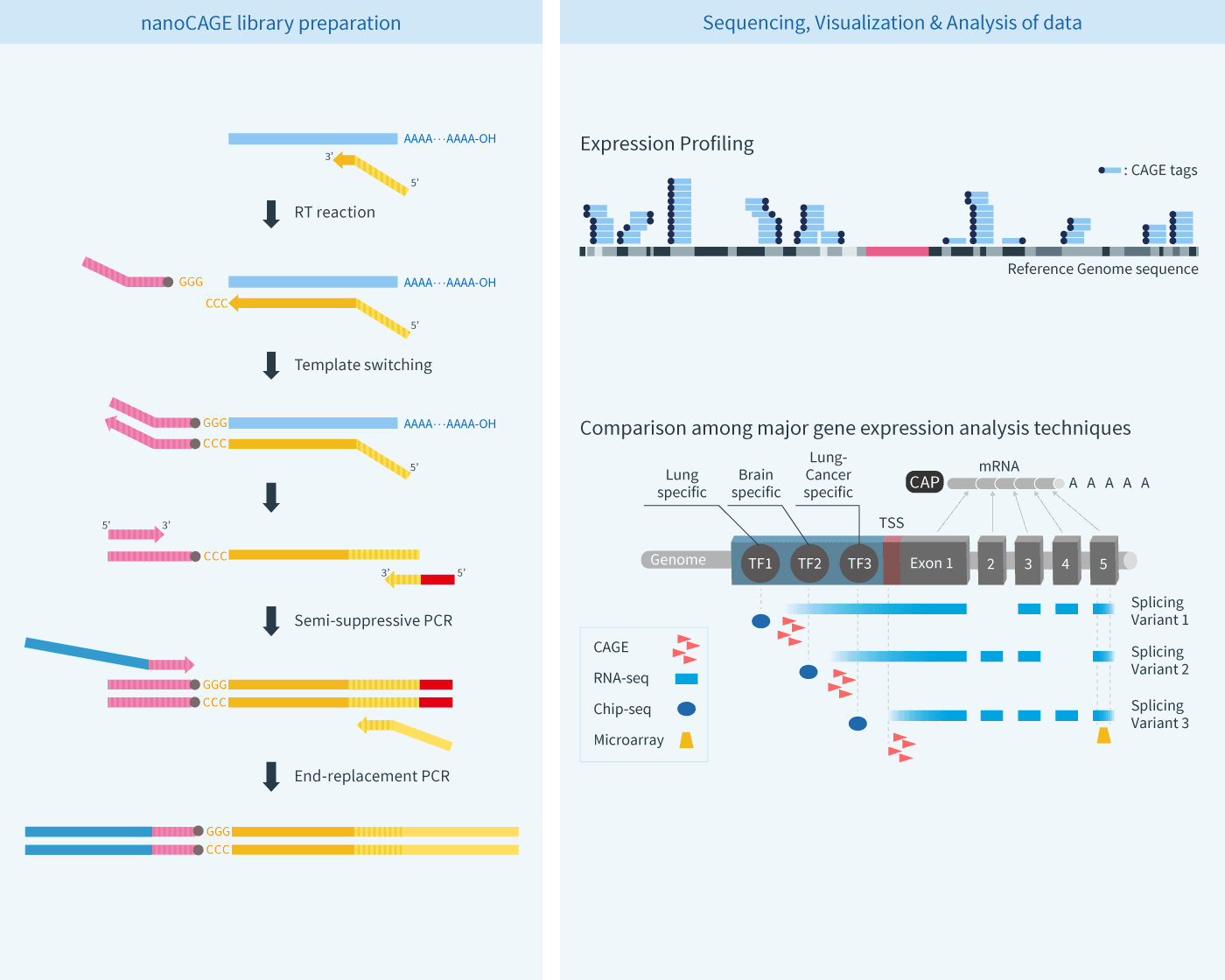
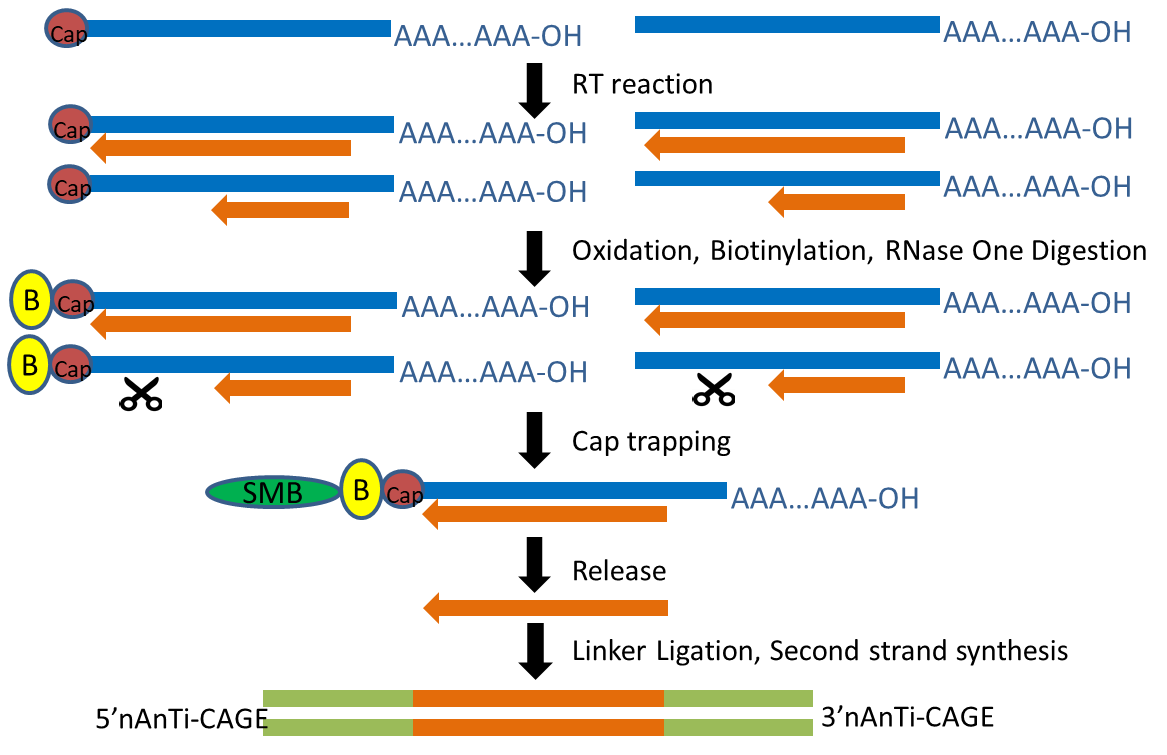
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター
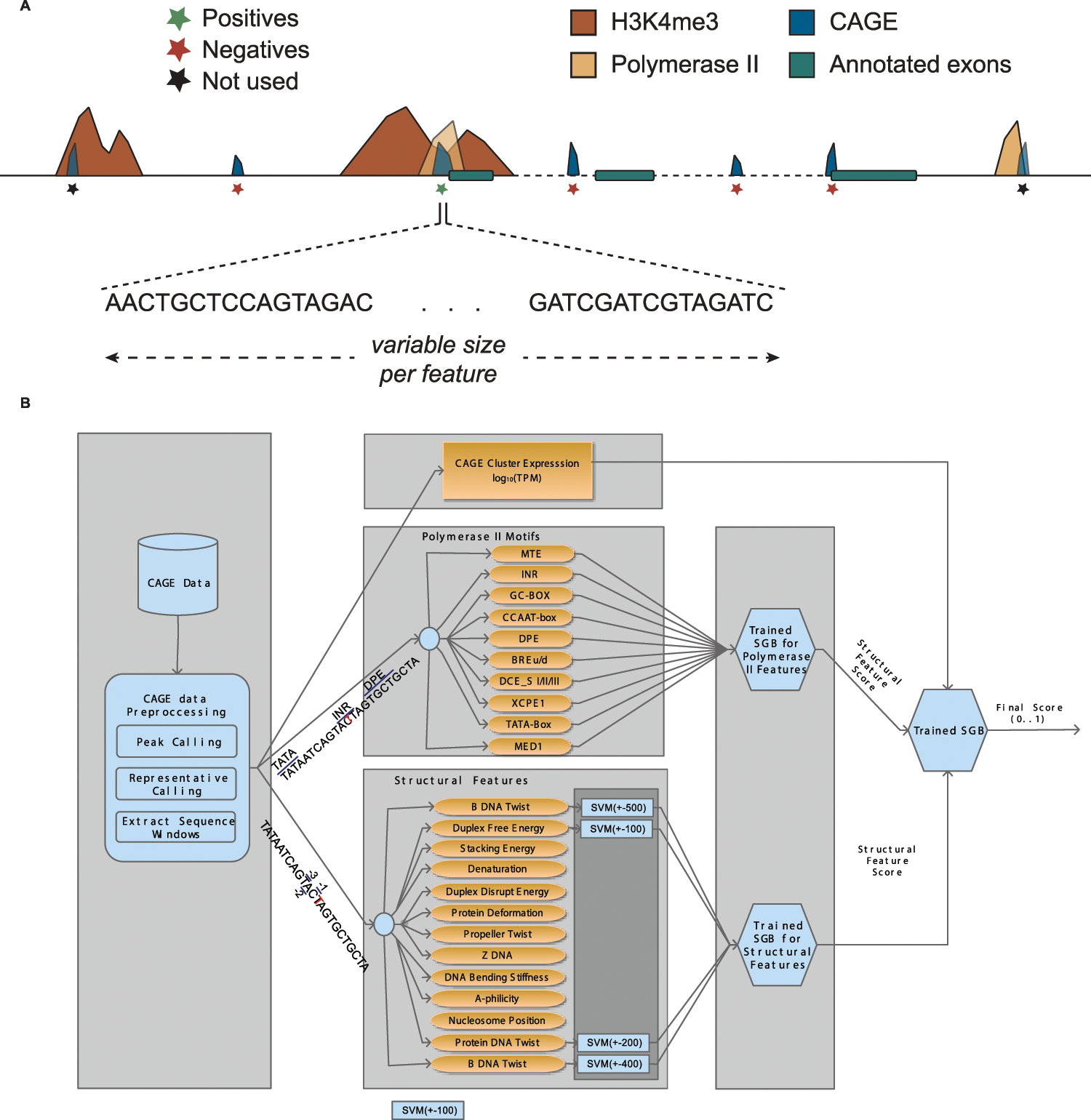
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports
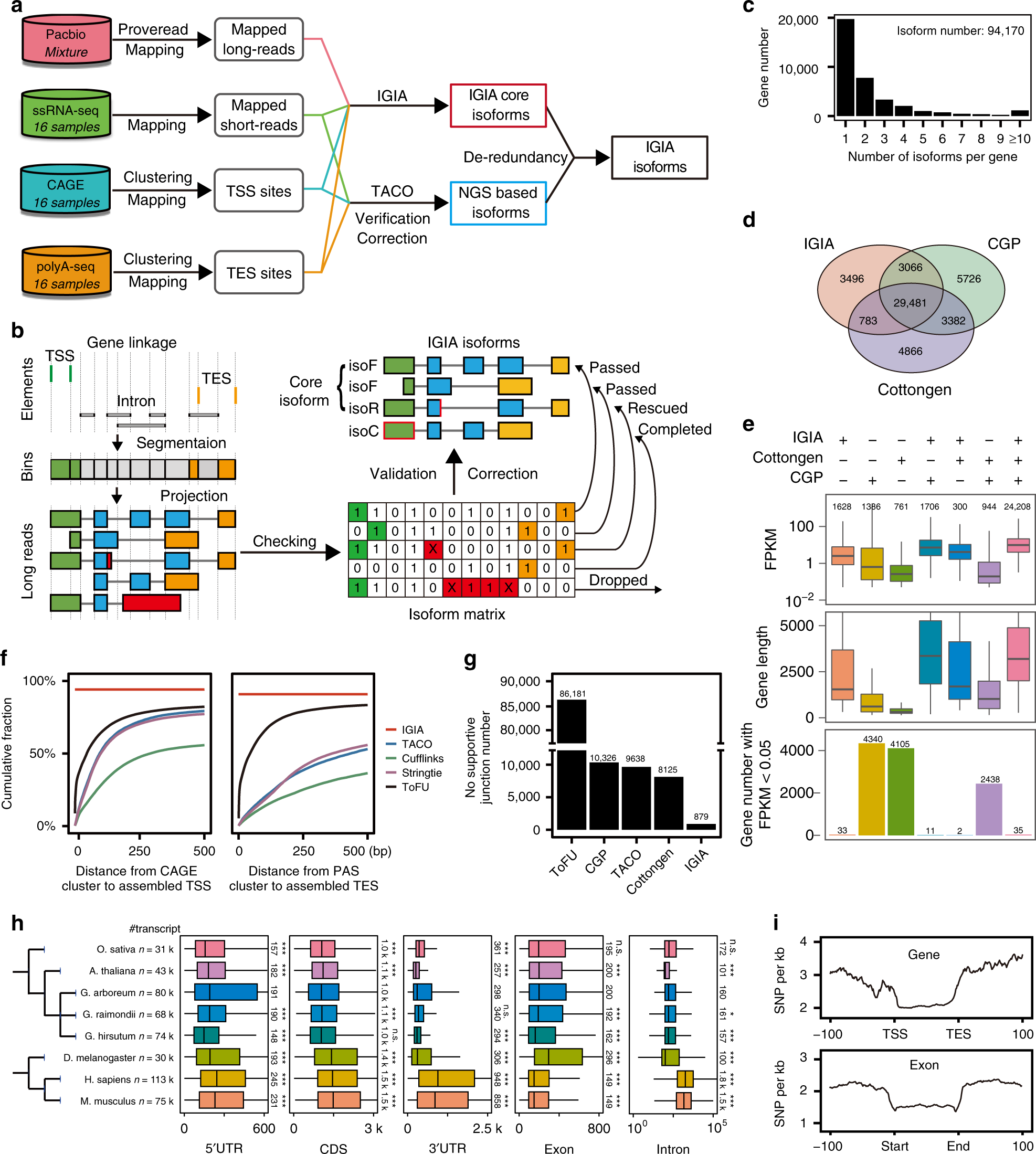
Multi-strategic RNA-seq analysis reveals a high-resolution transcriptional landscape in cotton | Nature Communications
CAGE-seq analysis of Epstein-Barr virus lytic gene transcription: 3 kinetic classes from 2 mechanisms | PLOS Pathogens

Cap analysis of gene expression (CAGE) sequencing reveals alternative promoter usage in complex disease | bioRxiv
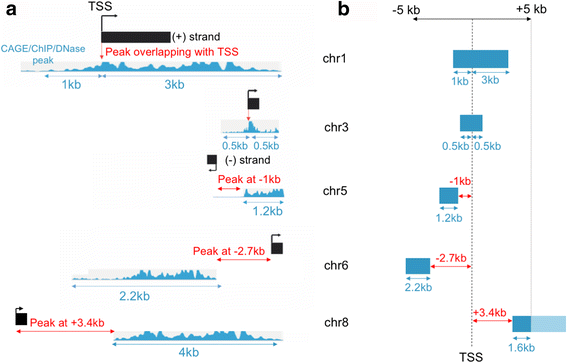
Zipper plot: visualizing transcriptional activity of genomic regions | BMC Bioinformatics | Full Text